URL
http://www.flyrnai.org/flyprimerbank
Evaluation of specific endogenous RNA transcript levels is important for understanding transcription regulation. It is also useful for independent confirmation of results obtained using techniques such as microarray analysis, RNA-seq and RNAi knockdown. We adapted and refined the algorithms used for the mammalian PrimerBank efforts for the model species Drosophila melanogaster and designed at least three primer pairs for each Drosophila gene. You can get primer information for fly genes one gene at a time or in batch mode. You can also view information like primer sequences, isoform specificity, location on transcripts, and any available validation results or/and user feedback here at FlyPrimerBank.
Looking for qPCR primers for transposable element detection? One primer design resource for TEs can be found in supplemental information provided in Czech et al. 2013 "A Transcriptome-wide RNAi Screen in the Drosophila Ovary Reveals Factors of the Germline piRNA Pathway" in Molecular Cell. DOI:https://doi.org/10.1016/j.molcel.2013.04.007
Tutorial Video
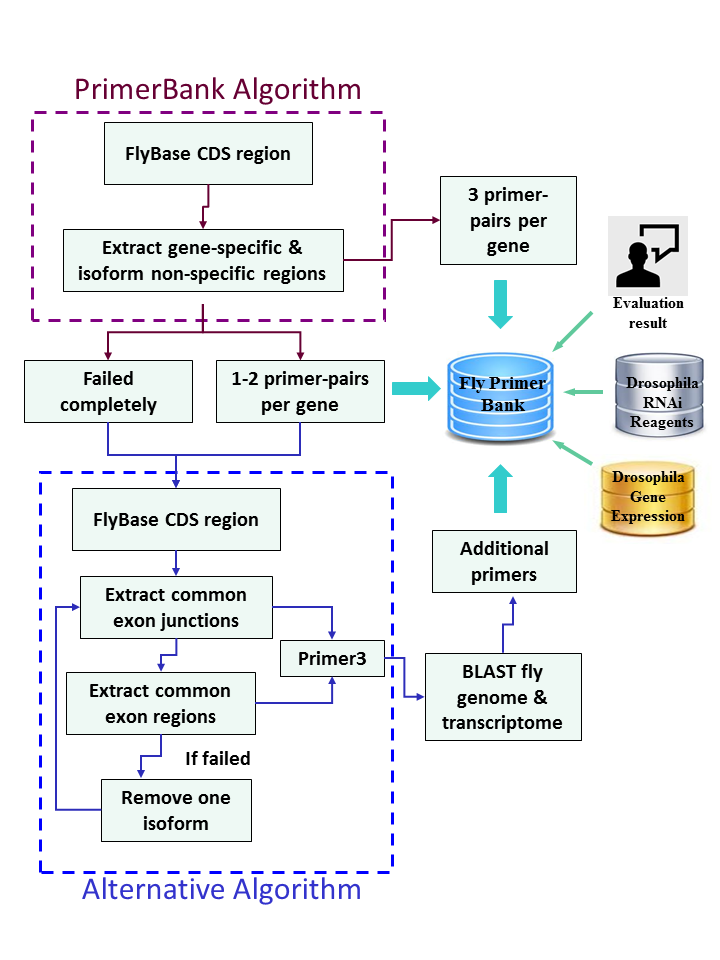
Workflow of FlyPrimerBank
The algorithm used for primer design was adopted from PrimerBank for Drosophila genes. For genes that failed design by this algorithm or do not have enough coverage, a different algorithm was used. Altogether, each gene is covered by at least 3 primer pairs at FlyPrimerBank.
PrimerBank workflow diagram
FlyPrimerBank Input Options
A list of identifiers for fly genes, including gene or protein identifiers like FBgn numbers, CG numbers and gene symbols.
For example,
dom
aay
FBgn0004066
CG18740
For example,
>wg-PA
ATGGATATCAGCTATATCTTCGTCATCTGCCTGATGGCCCTGTGCAGCGG
CGGCAGCAGTCTCAGCCAAGTCGAGGGCAAACAGAAATCCGGAAGGGGCC
GGGGCTCCATGTGGTGGGGCATTGCCAAGGTCGGCGAACCCAACAACATT
ACGCCCATCATGTACATGGACCCAGCGATCCACTCTACGTTGAGAAGGAA
ACAGCGACGCCTGGTCAGGGACAATCCCGGTGTACTGGGAGCCCTGGTCA
AGGGCGCCAACTTGGCCATTAGCGAGTGCCAACACCAGTTCAGAAATCGC
CGCTGGAACTGCTCGACGAGAAACTTCTCGAGGGGCAAAAATCTATTCGG
CAAAATCGTTGATCGAGGCTGCCGAGAGACGAGCTTCATTTACGCAATCA
CCAGCGCGGCGGTGACCCACTCGATTGCCAGGGCCTGCAGTGAAGGAACG
ATAGAGTCCTGCACCTGCGACTACAGCCACCAGTCGAGATCTCCACAAGC
>InR:exon9
CTTGCAAATCCATGGACATCAGGAACATGGTGTCGCACTTCAATCAGCTG
GAGAACTGCACGGTCATCGAGGGCTTCCTGCTGATCGATTTGATAAACGA
CGCCAGCCCTCTGAACAGAAGCTTTCCAAAACTGACCGAGGTCACAGATT
ATATCATAATCTACCGTGTGACTGGATTGCACTCGCTGTCAAAGATCTTT
CCCAATCTGAGCGTCATTAGGGGAAACAAGCTGTTCGACGGATATGCCTT
GGTCGTCTACTCGAATTTCGACCTCATGGATTTGGGACTTCACAAGCTAC
GATCCATAACCAGAGGCGGTGTGCGGATTGAGAAGAATCATAAGCTGTGC
TATGATAGGACCATCGATTGGCTGGAAATTCTGGCGGAAAACGAAACCCA
ACTGGTGGTGCTGACAGAGAACGGCAAGGAGAAGGAGTGCAGGCTTTCCA
AGTGCCCGGGGGAGATCAGAATTGAGGAGGGGCACGATACCACGGCTATT
GAGGGAGAGCTTAATGCCAGTTGTCAGCTGCACAATAATAGGCGCCTGTG
CTGGAACAGCAAACTCTGCCAGACGA
Overlap of PCR primers with RNAi reagents
To test knockdown following RNAi, you should avoid amplifying the reagent itself, and instead do the qPCR analysis on a different region of the transcript. This is especially relevant when the RNAi reagent is a long double-stranded RNA (dsRNA). To help with this, we included the predicted overlap of each predicted amplified sequence with dsRNAs from several public resources. The RNAi reagents for which we pre-computed these predictions include DRSC and VDRC dsRNAs for cell-based knockdown and long hairpin constructs for in vivo knockdown from the TRiP and NIG-Japan collections. For the RNAi reagents that are not pre-computed, you can evaluate the overlap by doing a pairwise BLAST between the qPCR amplicon and the RNAi reagent sequences.
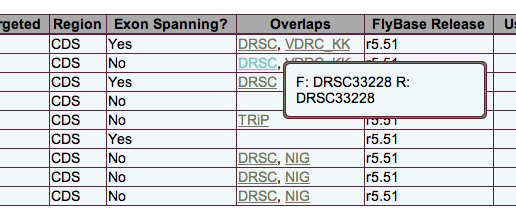
For each primer pair, the Overlaps column of the display lists any collections that include RNAi reagents which overlap with that pair. (Only those collections selected by the user are shown.) For the details, hover your mouse over the collection name. A tooltip will appear listing all reagents that overlap with the forward ("F:") and reverse ("R:") primers respectively. In the example above, a single reagent spans both primers and is therefore listed for both. The specific primer overlap is also indicated in the download file.
FlyBase data
Where possible primer data is based on FlyBase release r5.51 (May 2013). 164 primers do not map to FlyBase 5.51 and are therefore based on r5.44.
Expression data
Primers are less likely to work if the target genes are expressed at very low levels. We recommend using a tool such as CellExpressionData, DGET, or FlyBase views of modENCODE data to check expression of your target genes in the cell type(s), stage(s), and tissue(s) in which you will test them.
