Our DIOPT ortholog search tool has been updated to include the option to search for orthologs of a gene in all other species included. So you can search with, for example, a fly gene, and see orthologs in human, mouse, rat, frog, worm, and yeast.
Click on the button "show summary of top scores" to see a heat map view of the top-scoring ortholog matches in other species to your query species. This feature helps you see quickly if a gene has been conserved across many species or is, for example, only found in vertebrates.
As always, our tool supports batch-mode searches (you can enter more than one gene at a time) and allows you to download results as a file.
Feedback on DIOPT and our other online tools is welcome.
Click here for a direct link to the DIOPT ortholog search tool.

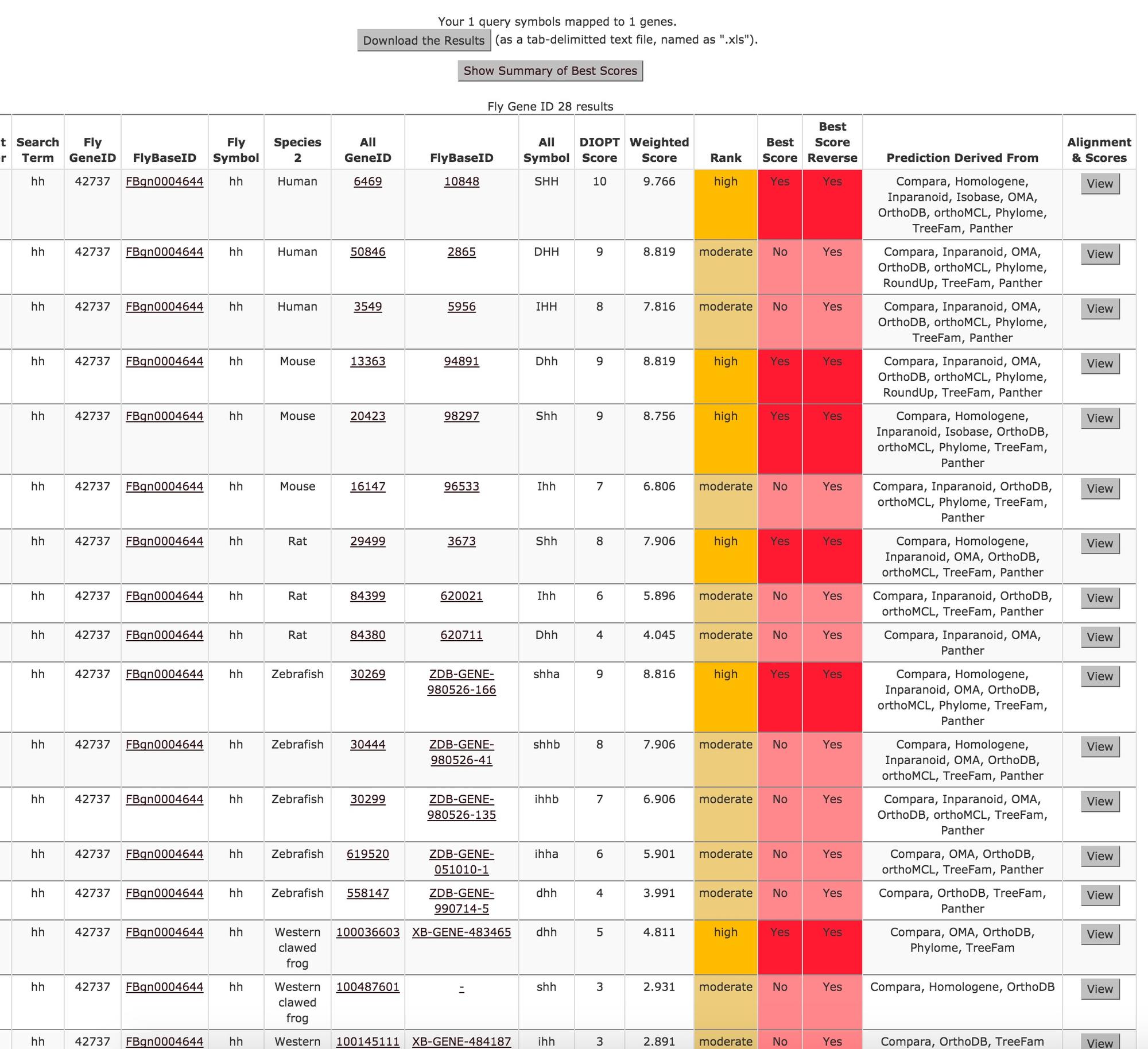
Results of an "all" search with Drosophila hh, wg, Atg1, and p53, followed by a click to view the "Summary of Best Scores" heatmap:
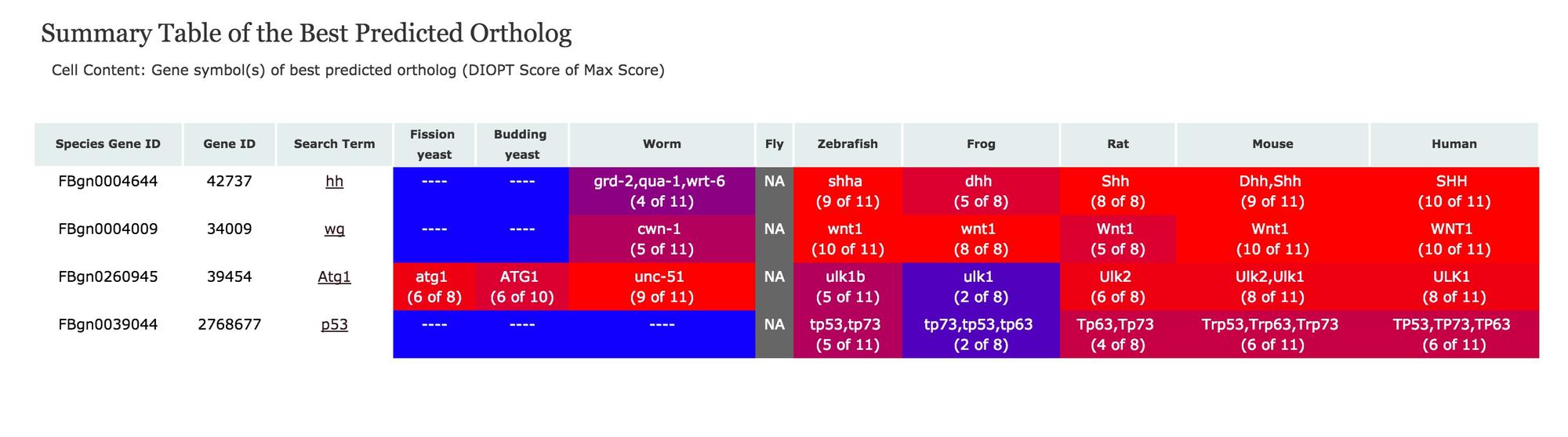
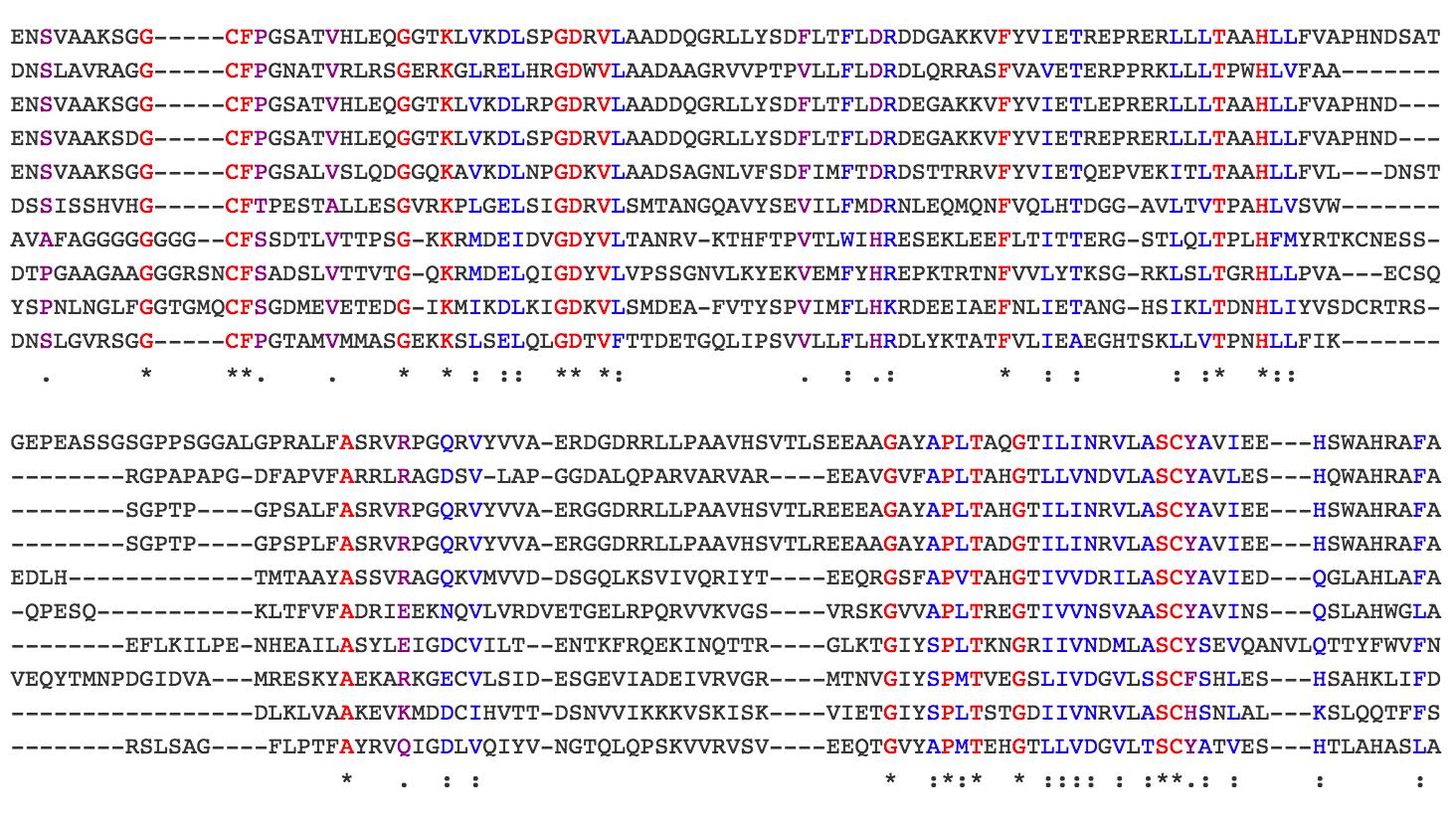
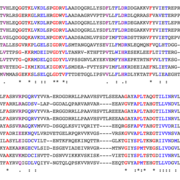
And from a best ortholog summary table, you can further link to a view of the domains present in each ortholog and multi-sequence alignment (see here for Hh orthologs). We are working on further improvements and welcome your feedback.